Week 1: Introduction to py-rcsb-api
Topics
- What
py-rcsb-apiis - Installation and imports
- Search API basics
- Data API basics
- Simple examples
- Hands-on exercises
- Resources
What is py-rcsb-api?
py-rcsb-api is the current Python toolkit for accessing RCSB PDB API services.
According to the official documentation, it provides Python interfaces for:
- the Search API
- the Data API
- the Sequence Coordinates API (not included in this workshop)
- the Model Server API (not included in this workshop)
For more information, visit:
Part 1: Search API
The Search API is used to find PDB entries that match search conditions. This is the easiest place to start.
Text search
A TextQuery works like a keyword search.
from rcsbapi.search import TextQuery
query = TextQuery(value="Hemoglobin")
results = list(query())
print(f"Found {len(results)} entries")
print(results[:10])
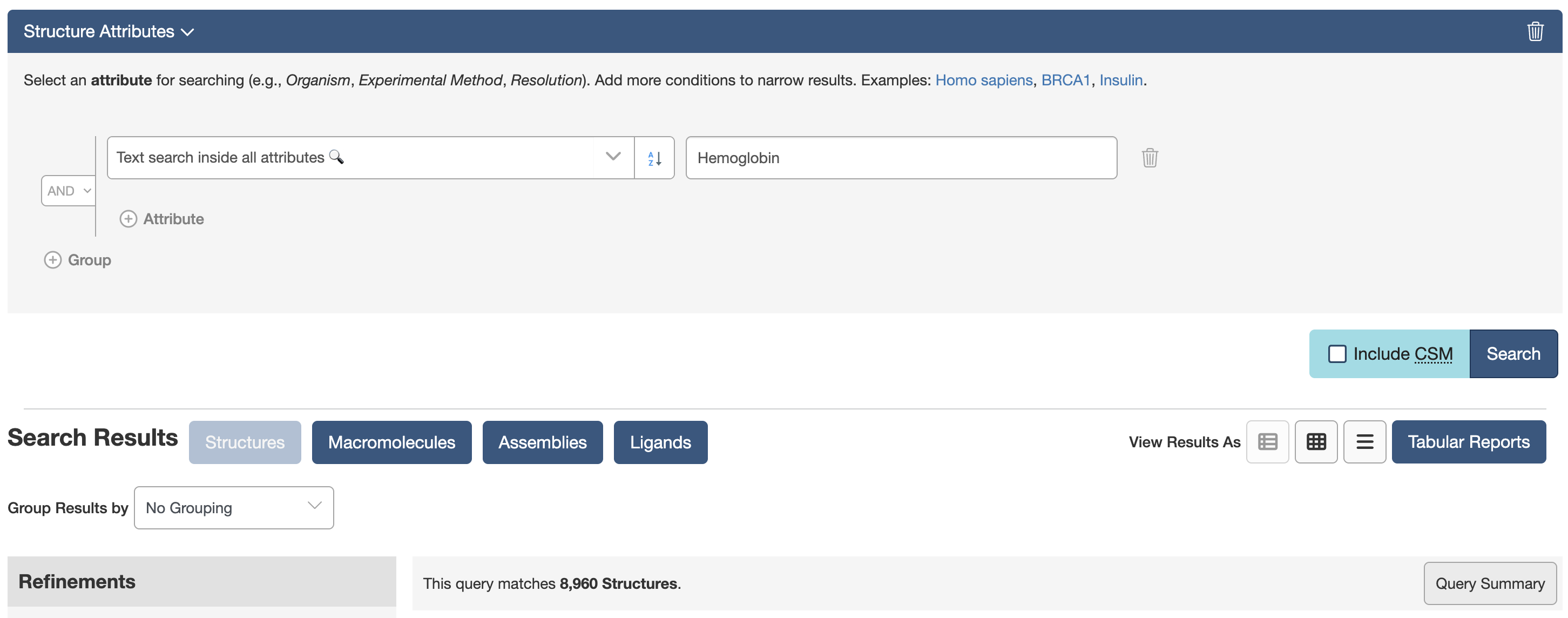
Attribute search
An AttributeQuery searches a specific field.
from rcsbapi.search import AttributeQuery
query = AttributeQuery(
attribute="rcsb_entity_source_organism.scientific_name",
operator="exact_match",
value="Homo sapiens"
)
results = list(query())
print(f"Found {len(results)} entries")
print(results[:10])
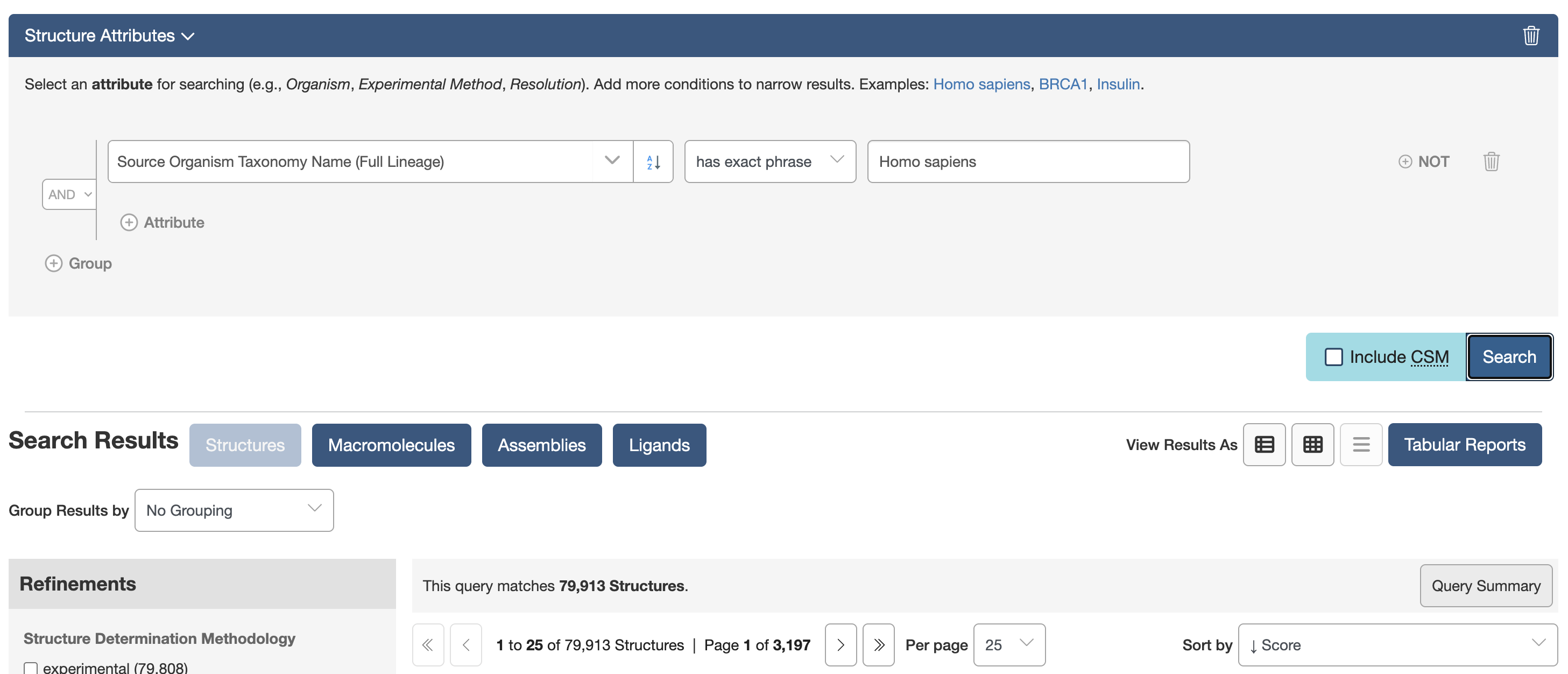
You can also use the shorter attribute syntax from search_attributes.
from rcsbapi.search import search_attributes as attrs
query = attrs.rcsb_entity_source_organism.scientific_name == "Homo sapiens"
results = list(query())
print(f"Found {len(results)} entries")
print(results[:10])
Combining queries
Queries can be combined with Python bitwise operators:
&means AND|means OR~means NOT
from rcsbapi.search import TextQuery
from rcsbapi.search import search_attributes as attrs
q1 = TextQuery(value="Hemoglobin")
q2 = attrs.rcsb_entity_source_organism.scientific_name == "Homo sapiens"
query = q1 & q2
results = list(query())
print(results[:10])
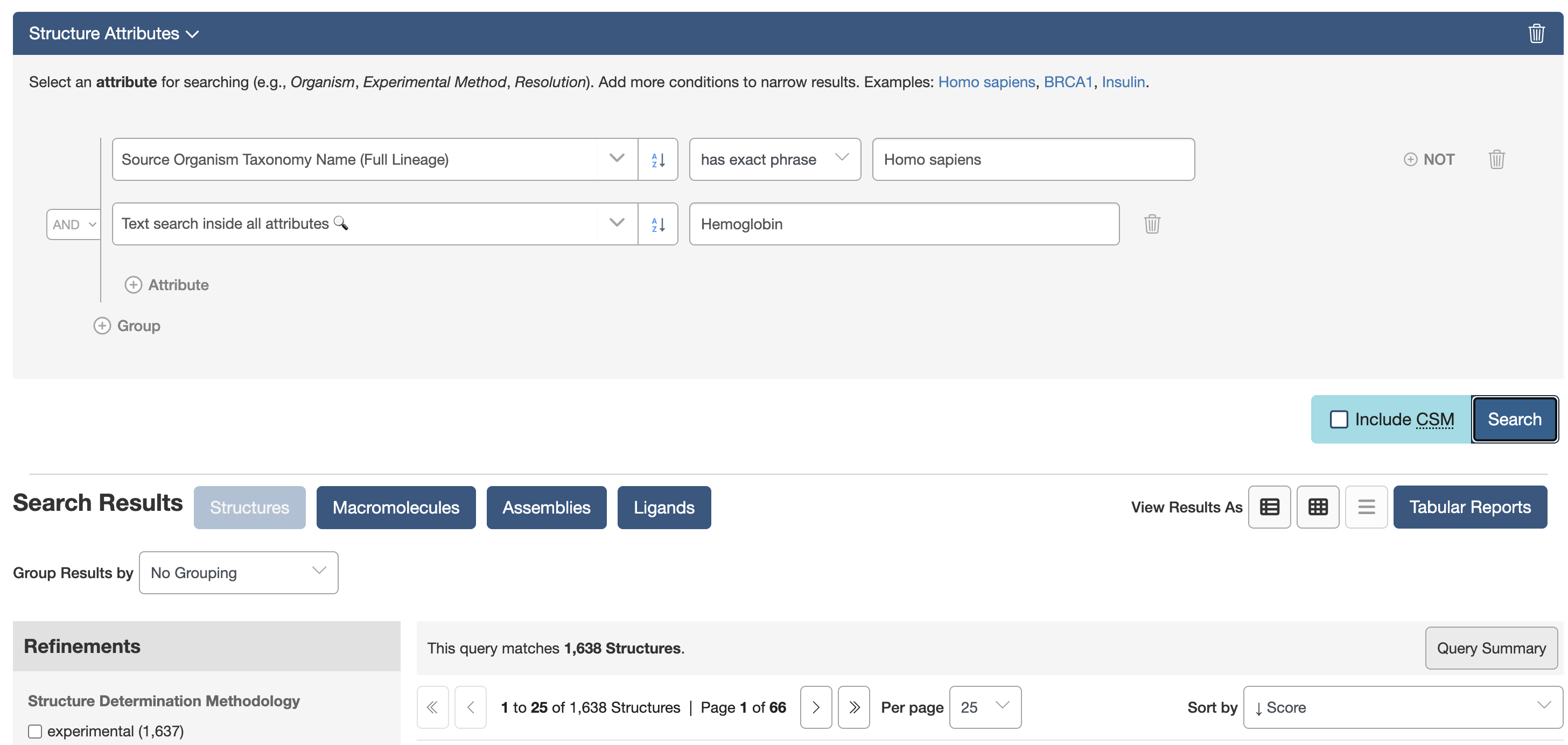
Another search example
q_method = attrs.exptl.method == "X-RAY DIFFRACTION"
q_resolution = attrs.rcsb_entry_info.resolution_combined < 2.5
query = q_method & q_resolution
results = list(query())
print(results[:10])
Useful references
Part 2: Data API
The Data API is used to retrieve structured information for specific IDs such as PDB entries. If the Search API answers the question “which structures match my conditions?”, the Data API answers the question “what details do these structures have?”
The official py-rcsb-api documentation explains that the Data API uses GraphQL under the hood.
In practice, this means:
- you start from an input object such as
entries - you provide one or more IDs, such as
4HHB - you request the exact fields you want back, such as
exptl.methodorstruct.title
DataQuery helps us build this GraphQL request in Python, so we do not have to write the GraphQL query by hand.
Basic pattern
A Data API query usually has three main parts:
input_typeThis says what kind of object you are querying, for exampleentries.input_idsThis is the list of IDs you want to look up.return_data_listThis is the list of fields you want to retrieve.
Example 1: get the experimental method for one structure
from rcsbapi.data import DataQuery
query = DataQuery(
input_type="entries",
input_ids=["4HHB"],
return_data_list=["exptl.method"]
)
result = query.exec()
print(result)
This asks for entry 4HHB and returns the value stored under exptl.method.
Example output
A simplified example of the returned result may look like this:
{
'data': {
'entries': [
{
'exptl': [
{'method': 'X-RAY DIFFRACTION'}
]
}
]
}
}
You usually need to inspect the result step by step to find the field you want.
Example 2: retrieve more than one field
from rcsbapi.data import DataQuery
query = DataQuery(
input_type="entries",
input_ids=["4HHB"],
return_data_list=["rcsb_id", "struct.title", "exptl.method"]
)
result = query.exec()
print(result)
Example 3: request data for multiple entries
This is useful when you already have a short list of PDB IDs from a search result.
from rcsbapi.data import DataQuery
query = DataQuery(
input_type="entries",
input_ids=["4HHB", "1CRN"],
return_data_list=["rcsb_id", "struct.title"]
)
result = query.exec()
print(result)
How the Data API relates to GraphQL
The RCSB Data API itself is based on GraphQL. GraphQL organizes data into:
- types: structured objects such as an entry
- fields: properties under those objects
- scalars: final values such as strings, numbers, or booleans
The py-rcsb-api package reads the schema and helps generate GraphQL automatically.
So instead of writing raw GraphQL like this:
{
entries(entry_ids: ["4HHB"]) {
exptl {
method
}
}
}
we can write the equivalent Python query:
from rcsbapi.data import DataQuery
query = DataQuery(
input_type="entries",
input_ids=["4HHB"],
return_data_list=["exptl.method"]
)
This is one of the main benefits of DataQuery.
It gives us a simpler Python interface, while still using the GraphQL Data API underneath.
Useful references
Hands-on exercises
Exercise 1
Use TextQuery to search for insulin and print the first 10 PDB IDs.
Exercise 2
Use search_attributes to find entries from Mus musculus and print the first 10 PDB IDs.
Exercise 3
Find entries determined by X-RAY DIFFRACTION with resolution better than 2.0 A.
Exercise 4
Search for kinase, then combine that query with a human organism filter.
Print the first 10 matching PDB IDs.
Exercise 5
Use 10 PDB IDs from Exercise 4 to find
- (1) number of biological assembly structures (bas),
- (2) number of chains in each structure,
- (3) corresponding UniProt ID.
- (4) Finally, please arrange the results in a dictionary as follow:
resulting_dictionary = { 'pdbID 1': { 'bas_1': { 'chain A': 'Uniprot ID 1', 'chain B': 'Uniprot ID 2', ... }, 'bas_2': {... } }, 'pdbID 2': {... }, ... }
One possible solution
In this exercise, we combine the Search API and the Data API. Instead of writing everything at once, it is easier to break the problem into smaller steps.
Step 1: use TextQuery and search_attributes to get 10 PDB IDs
from rcsbapi.search import TextQuery
from rcsbapi.search import search_attributes as attrs
q1 = TextQuery(value="kinase")
q2 = attrs.rcsb_entity_source_organism.scientific_name == "Homo sapiens"
query = q1 & q2
pdb_ids = list(query())[:10]
print(pdb_ids)
This gives us 10 PDB IDs from Exercise 4.
Step 2: use DataQuery(input_type="entries") to get assembly IDs and polymer entity IDs
For one PDB ID, we can ask the Data API for:
assembly_idspolymer_entity_ids
from rcsbapi.data import DataQuery
pdb_id = pdb_ids[0]
entry_query = DataQuery(
input_type="entries",
input_ids=[pdb_id],
return_data_list=[
"rcsb_entry_container_identifiers.assembly_ids",
"rcsb_entry_container_identifiers.polymer_entity_ids",
],
)
entry_result = entry_query.exec()
print(entry_result)
You can then extract the two lists like this:
entry_data = entry_result["data"]["entries"][0]
assembly_ids = entry_data["rcsb_entry_container_identifiers"].get("assembly_ids", [])
polymer_entity_ids = entry_data["rcsb_entry_container_identifiers"].get("polymer_entity_ids", [])
print("assembly_ids:", assembly_ids)
print("polymer_entity_ids:", polymer_entity_ids)
Step 3: use DataQuery(input_type="assemblies") to get the chain IDs in each biological assembly
For one biological assembly, the input ID format is PDBID-assembly_id.
For example:
assembly_query = DataQuery(
input_type="assemblies",
input_ids=[f"{pdb_id}-{assembly_ids[0]}"],
return_data_list=[
"rcsb_assembly_container_identifiers.assembly_id",
"pdbx_struct_assembly_gen.asym_id_list",
],
)
assembly_result = assembly_query.exec()
print(assembly_result)
To collect the chain IDs:
assembly_data = assembly_result["data"]["assemblies"][0]
assembly_gen = assembly_data.get("pdbx_struct_assembly_gen", [])
asym_ids_in_assembly = []
for item in assembly_gen:
asym_ids_in_assembly.extend(item.get("asym_id_list", []))
print(asym_ids_in_assembly)
Step 4: use DataQuery(input_type="polymer_entities") to map each chain to its UniProt ID
For one polymer entity, the input ID format is PDBID_entity_id.
For example:
polymer_entity_query = DataQuery(
input_type="polymer_entities",
input_ids=[f"{pdb_id}_{polymer_entity_ids[0]}"],
return_data_list=[
"rcsb_polymer_entity_container_identifiers.asym_ids",
"rcsb_polymer_entity_container_identifiers.uniprot_ids",
],
)
polymer_entity_result = polymer_entity_query.exec()
print(polymer_entity_result)
To build a chain-to-UniProt mapping:
polymer_entity_data = polymer_entity_result["data"]["polymer_entities"][0]
container = polymer_entity_data["rcsb_polymer_entity_container_identifiers"]
asym_ids = container.get("asym_ids", [])
uniprot_ids = container.get("uniprot_ids", [])
if len(uniprot_ids) == 0:
uniprot_value = None
elif len(uniprot_ids) == 1:
uniprot_value = uniprot_ids[0]
else:
uniprot_value = uniprot_ids
chain_to_uniprot = {}
for asym_id in asym_ids:
chain_to_uniprot[asym_id] = uniprot_value
print(chain_to_uniprot)
Step 5: arrange the result as a nested Python dictionary
The final target looks like this:
resulting_dictionary = {
"PDBID": {
"bas_1": {
"chain A": "UniProt ID",
"chain B": "UniProt ID",
}
}
}
To help with repeated chain IDs, we use a small helper function:
def unique_keep_order(items):
seen = set()
out = []
for x in items:
if x not in seen:
seen.add(x)
out.append(x)
return out
Final combined solution
from rcsbapi.search import TextQuery
from rcsbapi.search import search_attributes as attrs
from rcsbapi.data import DataQuery
def unique_keep_order(items):
seen = set()
out = []
for x in items:
if x not in seen:
seen.add(x)
out.append(x)
return out
# Step 1: repeat Exercise 4
q1 = TextQuery(value="kinase")
q2 = attrs.rcsb_entity_source_organism.scientific_name == "Homo sapiens"
query = q1 & q2
pdb_ids = list(query())[:10]
print("10 PDB IDs:")
print(pdb_ids)
# Step 2: build the final dictionary
resulting_dictionary = {}
for pdb_id in pdb_ids:
pdb_id = pdb_id.upper()
# Get assembly IDs and polymer entity IDs for this entry
entry_query = DataQuery(
input_type="entries",
input_ids=[pdb_id],
return_data_list=[
"rcsb_entry_container_identifiers.assembly_ids",
"rcsb_entry_container_identifiers.polymer_entity_ids",
],
)
entry_result = entry_query.exec()
entry_data = entry_result["data"]["entries"][0]
assembly_ids = entry_data["rcsb_entry_container_identifiers"].get("assembly_ids", [])
polymer_entity_ids = entry_data["rcsb_entry_container_identifiers"].get("polymer_entity_ids", [])
# Build chain -> UniProt mapping
chain_to_uniprot = {}
for entity_id in polymer_entity_ids:
polymer_entity_query = DataQuery(
input_type="polymer_entities",
input_ids=[f"{pdb_id}_{entity_id}"],
return_data_list=[
"rcsb_polymer_entity_container_identifiers.asym_ids",
"rcsb_polymer_entity_container_identifiers.uniprot_ids",
],
)
polymer_entity_result = polymer_entity_query.exec()
polymer_entity_data = polymer_entity_result["data"]["polymer_entities"][0]
container = polymer_entity_data["rcsb_polymer_entity_container_identifiers"]
asym_ids = container.get("asym_ids", [])
uniprot_ids = container.get("uniprot_ids", [])
if len(uniprot_ids) == 0:
uniprot_value = None
elif len(uniprot_ids) == 1:
uniprot_value = uniprot_ids[0]
else:
uniprot_value = uniprot_ids
for asym_id in asym_ids:
chain_to_uniprot[asym_id] = uniprot_value
# Build bas -> chain -> UniProt
resulting_dictionary[pdb_id] = {}
for assembly_id in assembly_ids:
assembly_query = DataQuery(
input_type="assemblies",
input_ids=[f"{pdb_id}-{assembly_id}"],
return_data_list=[
"rcsb_assembly_container_identifiers.assembly_id",
"pdbx_struct_assembly_gen.asym_id_list",
],
)
assembly_result = assembly_query.exec()
assembly_data = assembly_result["data"]["assemblies"][0]
assembly_gen = assembly_data.get("pdbx_struct_assembly_gen", [])
asym_ids_in_assembly = []
for item in assembly_gen:
asym_ids_in_assembly.extend(item.get("asym_id_list", []))
asym_ids_in_assembly = unique_keep_order(asym_ids_in_assembly)
bas_key = f"bas_{assembly_id}"
resulting_dictionary[pdb_id][bas_key] = {}
for asym_id in asym_ids_in_assembly:
if asym_id in chain_to_uniprot:
resulting_dictionary[pdb_id][bas_key][f"chain {asym_id}"] = chain_to_uniprot[asym_id]
print(resulting_dictionary)
Optional summary
If you also want to print the number of biological assemblies and the number of chains in each assembly:
for pdb_id, bas_dict in resulting_dictionary.items():
print(f"\n{pdb_id}")
print(f"Number of biological assemblies: {len(bas_dict)}")
for bas_name, chain_dict in bas_dict.items():
print(f" {bas_name}: {len(chain_dict)} chains")
Notes
- For
assemblies, the input ID format isPDBID-assembly_id, for example4HHB-1. - For
polymer_entities, the input ID format isPDBID_entity_id, for example4HHB_1. - Some chains may not have a UniProt mapping, so the value may be
None.